20.24.3. Eric is a commodity analyst who fit four different candidate AR(p) models to a series of oil prices. For each of the candidate models, he then retrieved the Akaike information criterion (AIC) and the Bayesian information criterion (BIC). The results are displayed below.
.. AIC and BIC values (table below) goes here …
As Eric interprets the results of these criteria (i.e., the AIC and BIC values), which of the following statements is TRUE?
- Both criteria (AIC and BIC) favor the AR(1) model
- Both criteria (AIC and BIC) favor the AR(2) model
- As both criteria are insignificant, any AR(p) model is a bad fit
- The criteria disagree and therefore the AIC’s preference should prevail because it prefers fewer lags
library(tidyverse)
library(scales)
library(gridExtra)
library(forecast)
library(tseries)
library(fpp2)
library(gt)
library(astsa)
# 175
set.seed(175)
ar2 <- arima.sim(model=list(order=c(2,0,0),ar = c(1.5,-.75)),n = 500)
ma2 <- arima.sim(model=list(order=c(0,0,2),ma = c(1.5,-.75)),n = 500)
ar1_fit <- sarima(ar2, p = 1, d = 0, q = 0)
initial value 1.185288
iter 2 value 0.473121
iter 3 value 0.473113
iter 4 value 0.473112
iter 5 value 0.473108
iter 6 value 0.473099
iter 7 value 0.473095
iter 8 value 0.473094
iter 9 value 0.473092
iter 10 value 0.473087
iter 11 value 0.473080
iter 12 value 0.473076
iter 13 value 0.473075
iter 14 value 0.473075
iter 15 value 0.473075
iter 16 value 0.473074
iter 17 value 0.473073
iter 18 value 0.473073
iter 19 value 0.473073
iter 20 value 0.473073
iter 21 value 0.473072
iter 22 value 0.473072
iter 23 value 0.473072
iter 24 value 0.473072
iter 25 value 0.473072
iter 26 value 0.473072
iter 27 value 0.473072
iter 27 value 0.473072
iter 27 value 0.473072
final value 0.473072
converged
initial value 0.475632
iter 2 value 0.475627
iter 3 value 0.475615
iter 4 value 0.475613
iter 5 value 0.475597
iter 6 value 0.475583
iter 7 value 0.475579
iter 8 value 0.475574
iter 9 value 0.475573
iter 10 value 0.475570
iter 11 value 0.475569
iter 12 value 0.475569
iter 13 value 0.475569
iter 13 value 0.475569
iter 13 value 0.475569
final value 0.475569
converged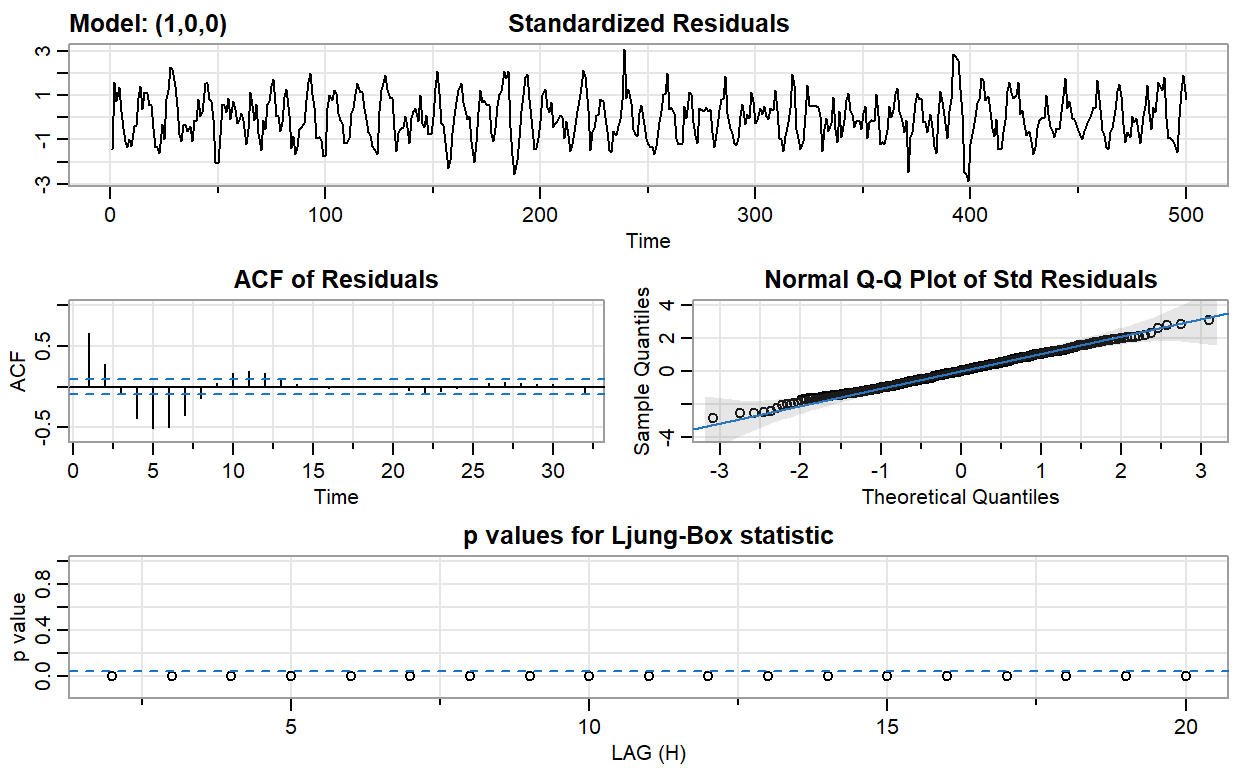
ar2_fit <- sarima(ar2, p = 2, d = 0, q = 0)
initial value 1.186082
iter 2 value 1.063752
iter 3 value 0.611728
iter 4 value 0.360300
iter 5 value 0.180452
iter 6 value 0.026222
iter 7 value 0.023222
iter 8 value 0.022348
iter 9 value 0.022310
iter 10 value 0.022284
iter 11 value 0.022258
iter 12 value 0.022258
iter 13 value 0.022258
iter 14 value 0.022258
iter 15 value 0.022258
iter 16 value 0.022258
iter 17 value 0.022258
iter 17 value 0.022258
iter 17 value 0.022258
final value 0.022258
converged
initial value 0.028108
iter 2 value 0.028095
iter 3 value 0.028087
iter 4 value 0.028086
iter 5 value 0.028086
iter 6 value 0.028085
iter 7 value 0.028085
iter 8 value 0.028085
iter 8 value 0.028085
final value 0.028085
converged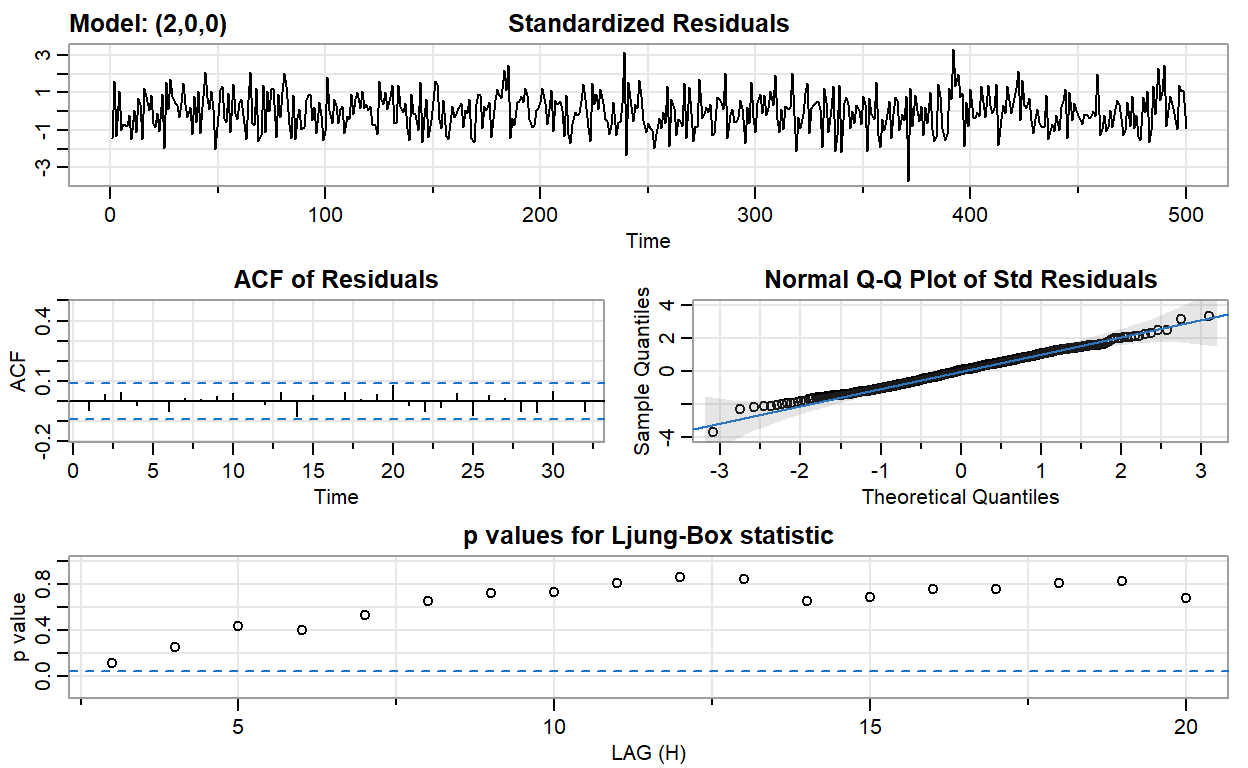
ar3_fit <- sarima(ar2, p = 3, d = 0, q = 0)
initial value 1.187086
iter 2 value 0.966297
iter 3 value 0.720122
iter 4 value 0.651535
iter 5 value 0.270365
iter 6 value 0.157749
iter 7 value 0.106223
iter 8 value 0.068752
iter 9 value 0.027592
iter 10 value 0.020453
iter 11 value 0.020449
iter 12 value 0.020445
iter 13 value 0.020430
iter 14 value 0.020418
iter 15 value 0.020418
iter 16 value 0.020418
iter 17 value 0.020418
iter 18 value 0.020418
iter 19 value 0.020418
iter 20 value 0.020418
iter 20 value 0.020418
iter 20 value 0.020418
final value 0.020418
converged
initial value 0.027093
iter 2 value 0.027090
iter 3 value 0.027084
iter 4 value 0.027079
iter 5 value 0.027079
iter 6 value 0.027078
iter 7 value 0.027078
iter 8 value 0.027078
iter 8 value 0.027078
iter 8 value 0.027078
final value 0.027078
converged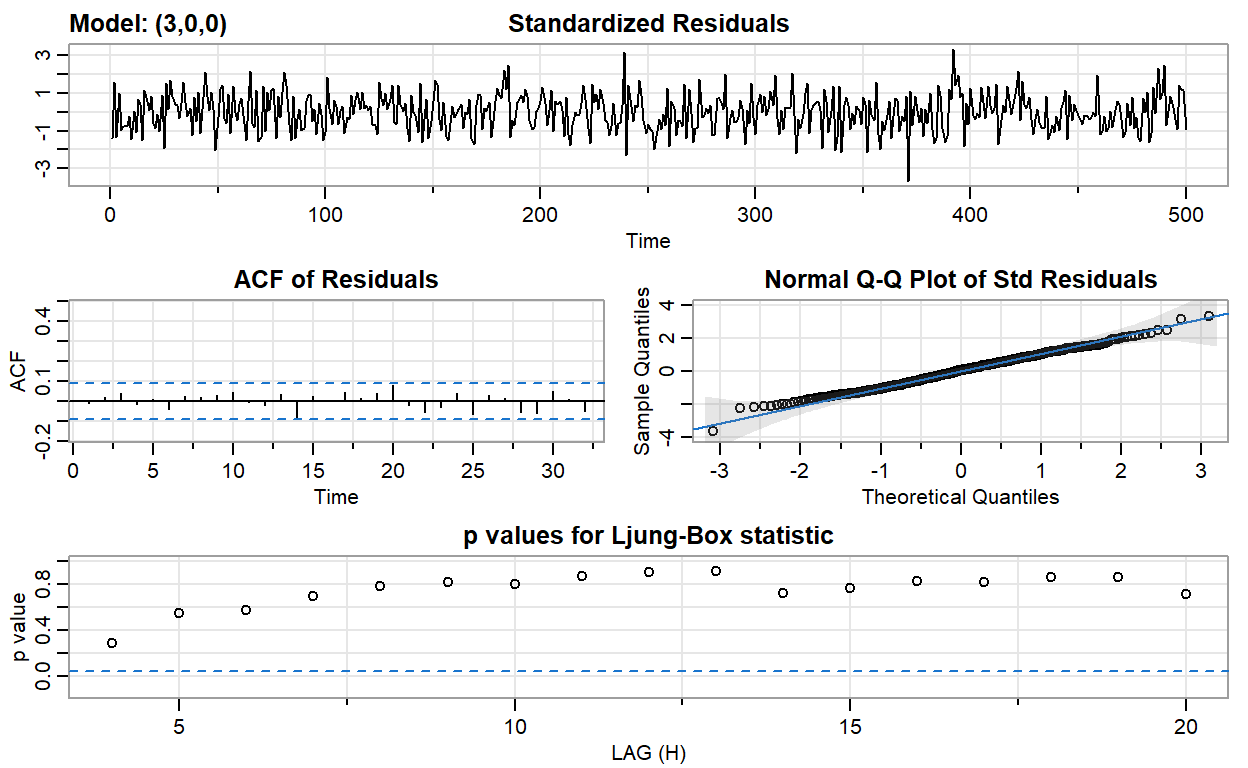
ar4_fit <- sarima(ar2, p = 4, d = 0, q = 0)
initial value 1.187677
iter 2 value 1.182760
iter 3 value 0.389837
iter 4 value 0.312734
iter 5 value 0.257361
iter 6 value 0.134886
iter 7 value 0.042493
iter 8 value 0.035479
iter 9 value 0.028843
iter 10 value 0.020356
iter 11 value 0.020308
iter 12 value 0.020305
iter 13 value 0.020305
iter 14 value 0.020305
iter 15 value 0.020304
iter 16 value 0.020304
iter 17 value 0.020304
iter 18 value 0.020304
iter 19 value 0.020304
iter 20 value 0.020304
iter 21 value 0.020304
iter 22 value 0.020304
iter 22 value 0.020304
iter 22 value 0.020304
final value 0.020304
converged
initial value 0.026957
iter 2 value 0.026941
iter 3 value 0.026940
iter 4 value 0.026938
iter 5 value 0.026937
iter 6 value 0.026936
iter 7 value 0.026936
iter 8 value 0.026936
iter 9 value 0.026936
iter 9 value 0.026936
iter 9 value 0.026936
final value 0.026936
converged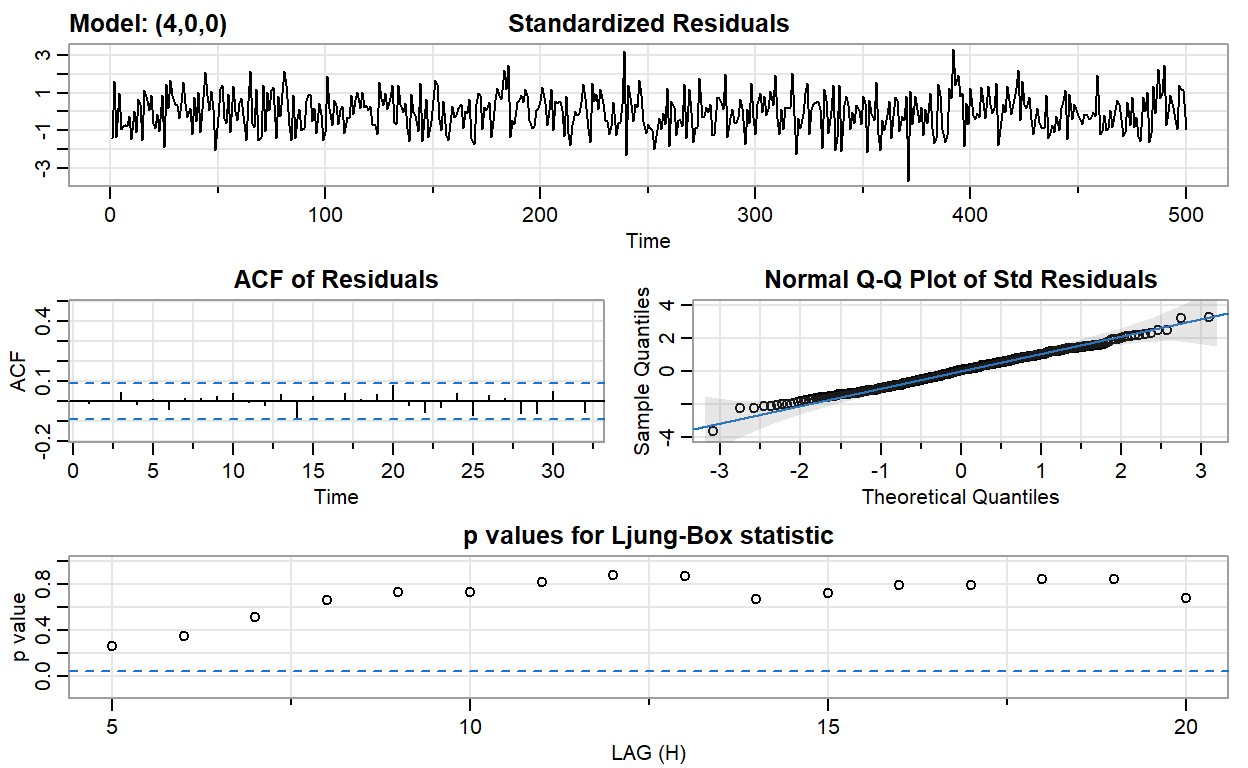
ma1_fit <- sarima(ma2, p = 0, d = 0, q = 1)
initial value 0.620720
iter 2 value 0.616967
iter 3 value 0.616503
iter 4 value 0.616503
iter 4 value 0.616503
iter 4 value 0.616503
final value 0.616503
converged
initial value 0.616515
iter 2 value 0.616515
iter 2 value 0.616515
iter 2 value 0.616515
final value 0.616515
converged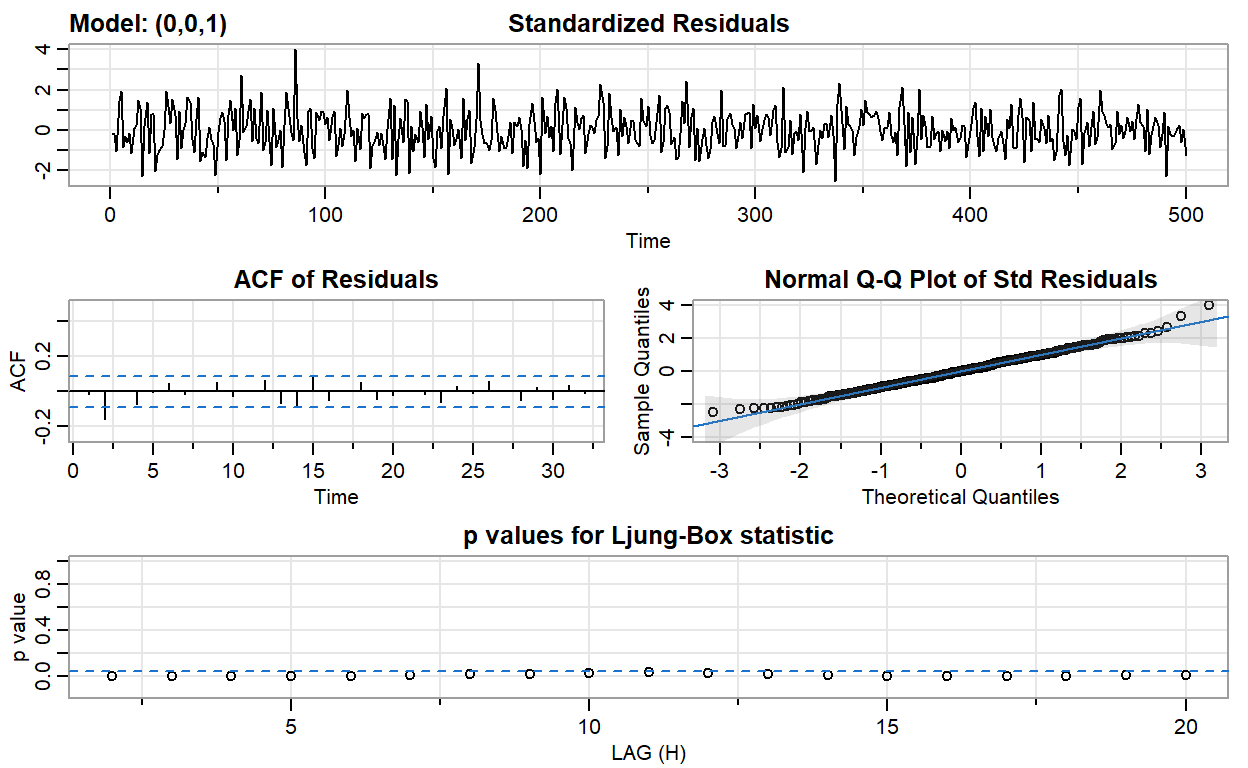
ma2_fit <- sarima(ma2, p = 0, d = 0, q = 2)
initial value 0.620720
iter 2 value 0.601563
iter 3 value 0.601097
iter 4 value 0.601053
iter 5 value 0.601053
iter 5 value 0.601053
iter 5 value 0.601053
final value 0.601053
converged
initial value 0.601132
iter 2 value 0.601132
iter 2 value 0.601132
iter 2 value 0.601132
final value 0.601132
converged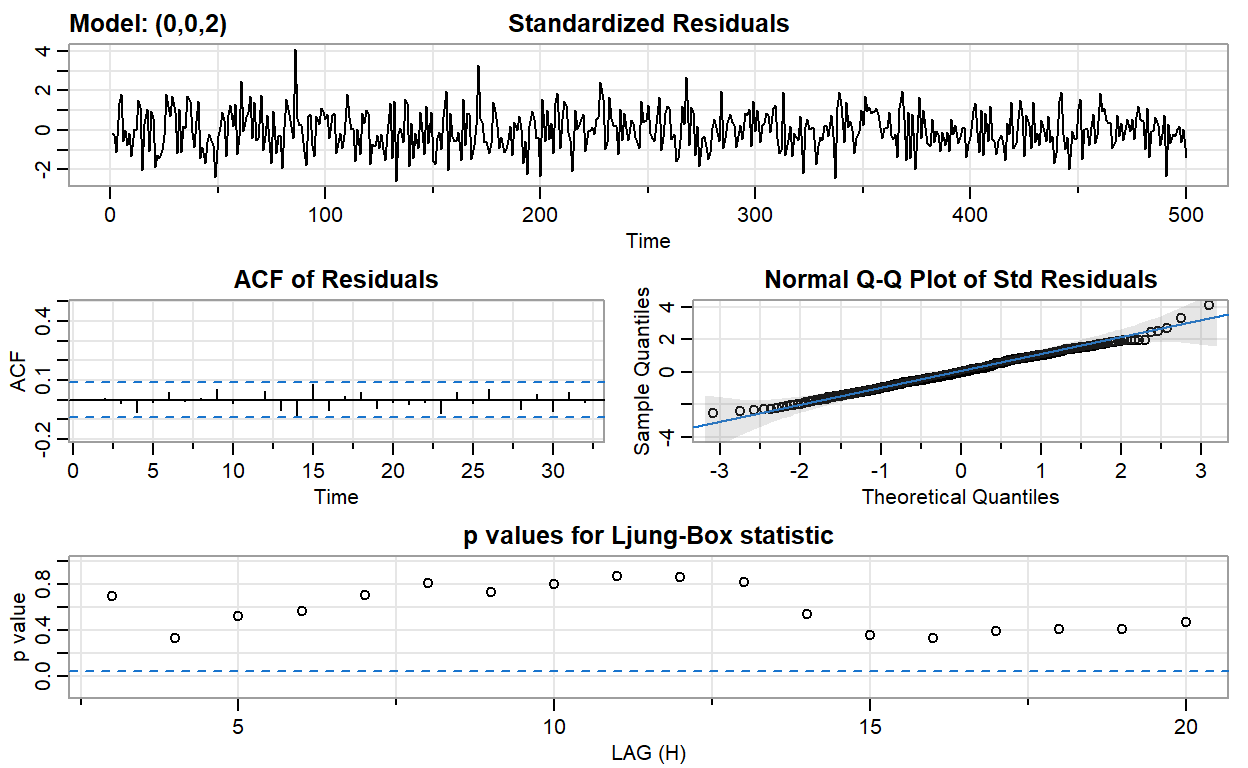
ma3_fit <- sarima(ma2, p = 0, d = 0, q = 3)
initial value 0.620720
iter 2 value 0.601584
iter 3 value 0.601080
iter 4 value 0.600978
iter 5 value 0.600977
iter 5 value 0.600977
iter 5 value 0.600977
final value 0.600977
converged
initial value 0.601058
iter 2 value 0.601058
iter 3 value 0.601058
iter 3 value 0.601058
iter 3 value 0.601058
final value 0.601058
converged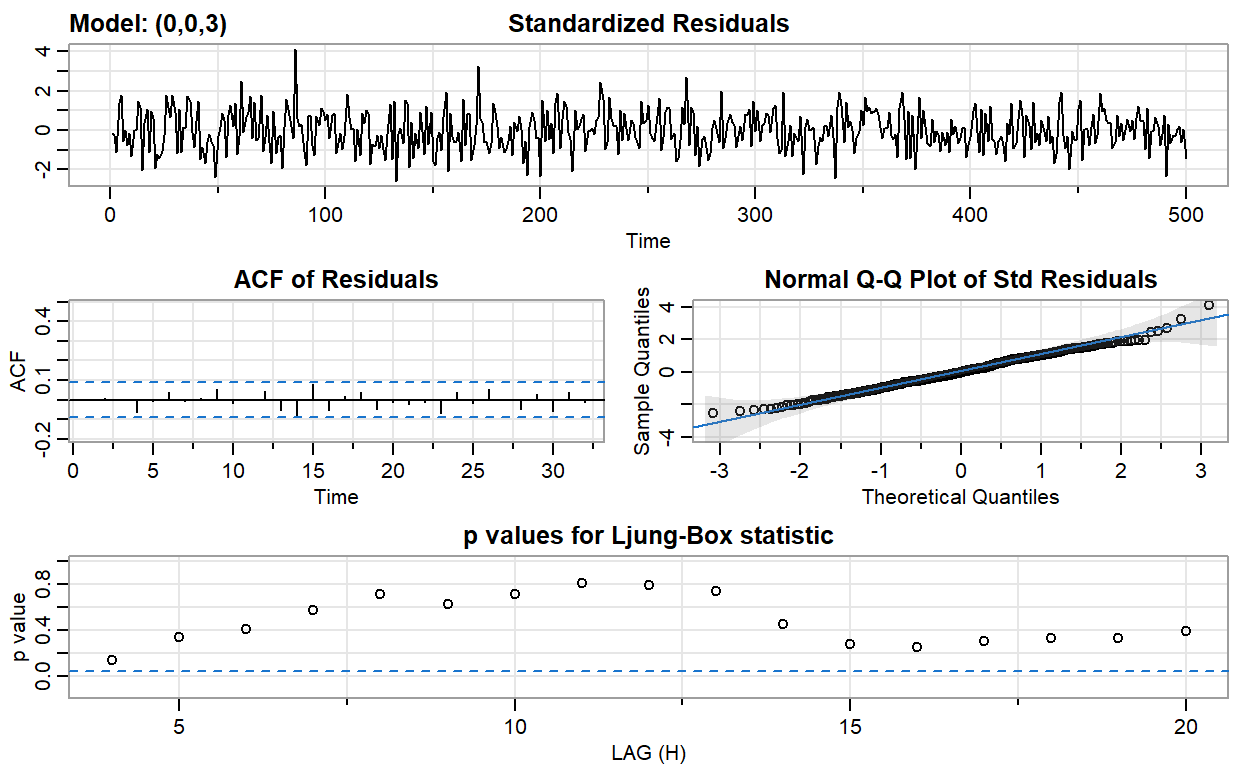
ma4_fit <- sarima(ma2, p = 0, d = 0, q = 4)
initial value 0.620720
iter 2 value 0.599626
iter 3 value 0.599377
iter 4 value 0.599342
iter 5 value 0.599340
iter 6 value 0.599340
iter 7 value 0.599340
iter 7 value 0.599340
iter 7 value 0.599340
final value 0.599340
converged
initial value 0.599424
iter 2 value 0.599424
iter 3 value 0.599424
iter 3 value 0.599424
iter 3 value 0.599424
final value 0.599424
converged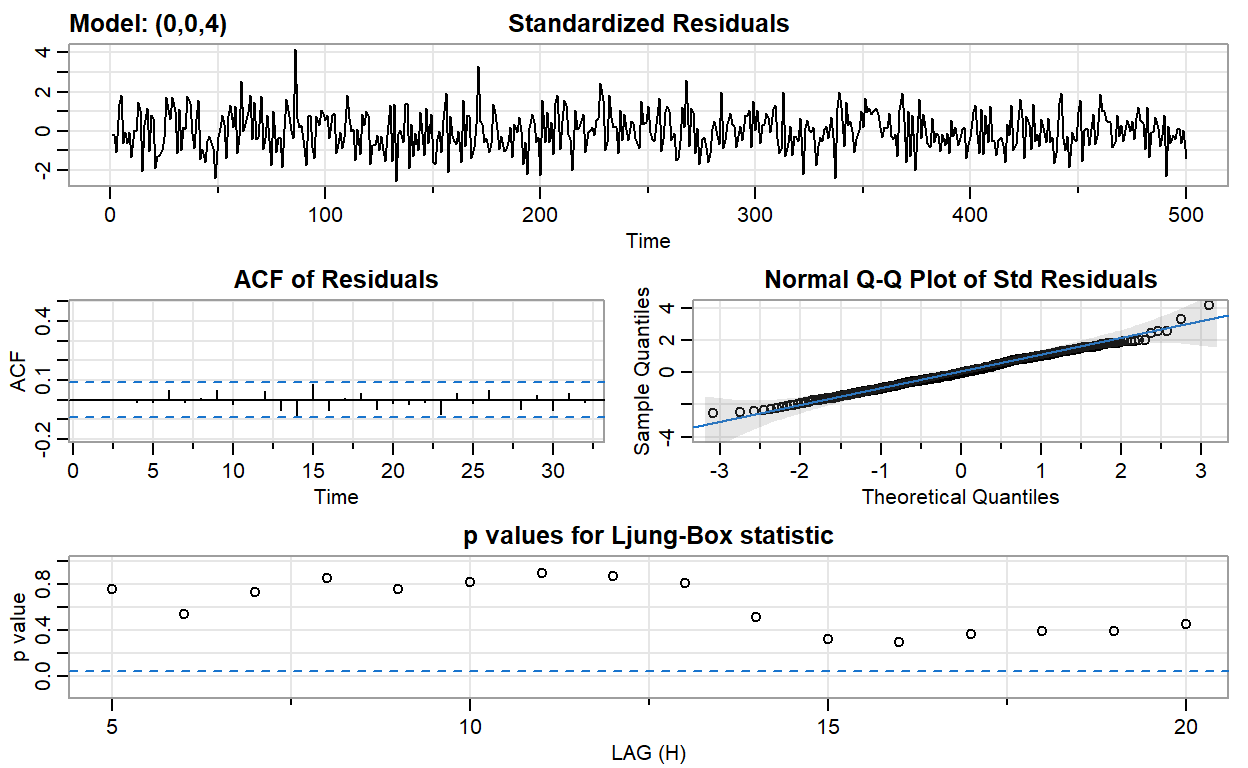
print("AIC")
[1] "AIC"ar1_fit$AIC
[1] 3.801016ar2_fit$AIC
[1] 2.910047ar3_fit$AIC
[1] 2.912034ar4_fit$AIC
[1] 2.915749print("BIC")
[1] "BIC"ar1_fit$BIC
[1] 3.826304ar2_fit$BIC
[1] 2.943764ar3_fit$BIC
[1] 2.95418ar4_fit$BIC
[1] 2.966324AIC_ar2 <- c(ar1_fit$AIC, ar2_fit$AIC, ar3_fit$AIC, ar4_fit$AIC)
BIC_ar2 <- c(ar1_fit$BIC, ar2_fit$BIC, ar3_fit$BIC, ar4_fit$BIC)
AIC_BIC_ar2 <- cbind(AIC_ar2, BIC_ar2) %>% data.frame()
models <- c("AR(1)", "AR(2)", "AR(3)", "AR(4)")
models_AIC_BIC <- cbind(models, AIC_BIC_ar2)
models_AIC_BIC_gt <- gt(models_AIC_BIC)
models_AIC_BIC_gt <-
models_AIC_BIC_gt %>%
tab_options(
table.font.size = 14
) %>% tab_style(
style = cell_text(weight = "bold"),
locations = cells_body()
# ) %>% tab_style(
# style = cell_text(color = "cadetblue"),
# locations = cells_column_labels(
# columns = vars(model, 'h (lags)', statistic, 'p-value')
# )
) %>% tab_header(
title = md("**AIC and BIC values**"),
subtitle = "Four candidate models: AR(1) ... AR(4)"
) %>% fmt_number(
columns = vars(AIC_ar2, BIC_ar2),
decimals = 5
) %>% tab_source_note(
source_note = md("AIC = Akaike information criterion")
) %>% tab_source_note(
source_note = md("BIC = Bayesian information criterion")
# ) %>% cols_width(
# vars(model) ~ px(70),
# vars(statistic, 'p-value') ~ px(90)
) %>% cols_label (
models = md("**model**"),
AIC_ar2 = md("**AIC**"),
BIC_ar2 = md("**BIC**")
) %>% tab_options(
heading.title.font.size = 16,
heading.subtitle.font.size = 14
)
models_AIC_BIC_gt
| AIC and BIC values | ||
|---|---|---|
| Four candidate models: AR(1) ... AR(4) | ||
| model | AIC | BIC |
| AR(1) | 3.80102 | 3.82630 |
| AR(2) | 2.91005 | 2.94376 |
| AR(3) | 2.91203 | 2.95418 |
| AR(4) | 2.91575 | 2.96632 |
| AIC = Akaike information criterion | ||
| BIC = Bayesian information criterion | ||